We work at the spatial biology-technology interface to understand chromatin<–>synapse communication in the mammalian brain.
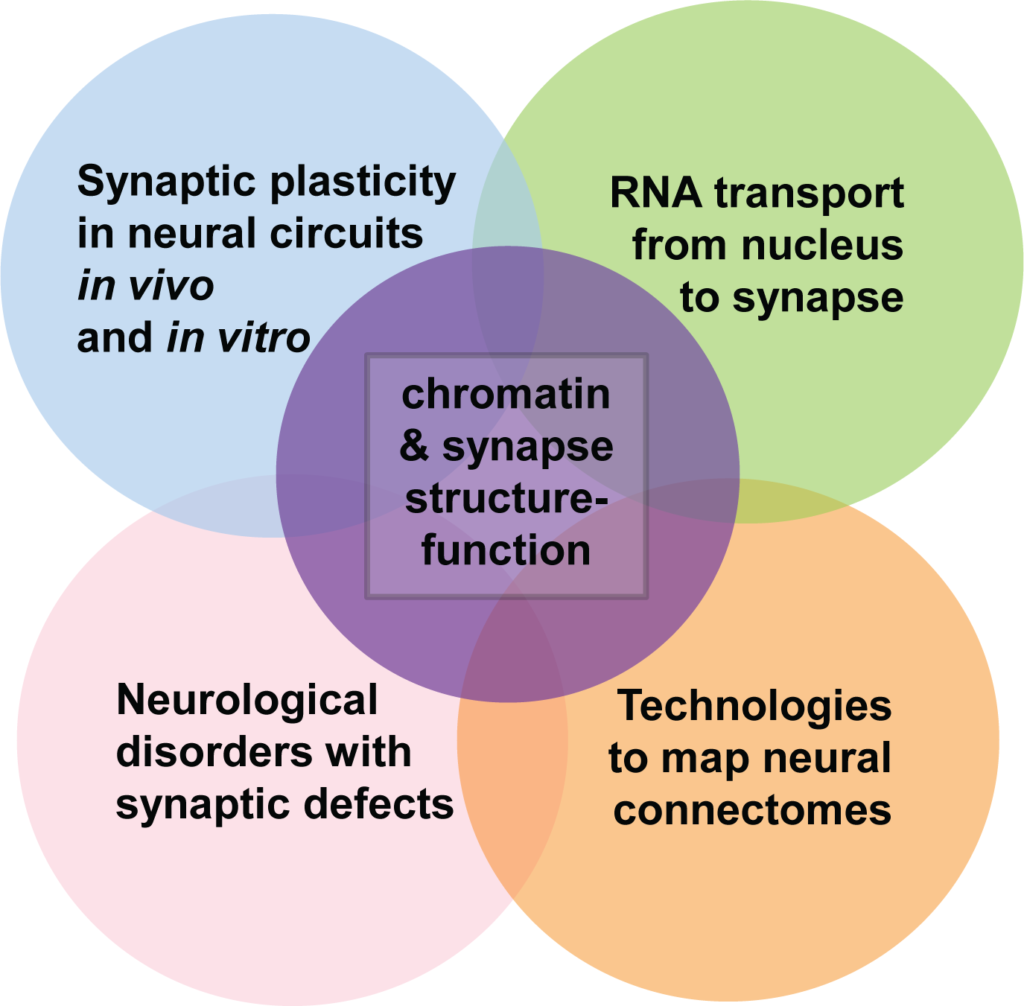
The Cremins Laboratory works at the spatial biology-technology interface to investigate the structure-function relationship of connections across the scales of chromatin, synapses, and circuits in the developing and diseased mammalian brain. Neurons exhibit elaborate structures at multiple scales, including diverse long-range axonal projections to form circuits, a range of complex dendritic spine geometries at the synapse, and the folding of the 10 nm chromatin fiber into compartments, topologically associating domains (TADs), and long-range looping interactions in the nucleus.
We have thus far focused on the nucleus to elucidate chromatin’s higher-order folding patterns at kilobase-resolution during neural lineage commitment, maturation, activity stimulation, and in neurological disorders. In mammals, the genetic sequence is >2 meters when stretched out end-to-end. It is folded into exquisite structures to fit into a nucleus roughly the size of a pin’s head. At the lab’s inception, it remained unclear how genomes are folded in a cell type-specific manner below the resolution of a Megabase, and whether and how higher-order folding could deterministically influence genome function. Our major contribution has been to provide early foundational insights into the genome’s structure-function relationship. We have elucidated how chromatin works through long-range mechanisms to govern the establishment of new gene expression, repeat instability, and replication origin firing patterns when mammalian cells transition states in the healthy and diseased brain.
Overall, by adding a spatial, third dimension to our understanding of chromatin’s regulatory mechanisms, the Cremins lab has uncovered foundational basic insights into the genome’s structure-function relationship in the mammalian brain and in neurological disorders. Looking forward, the lab will focus on mechanisms underlying chromatin-synapse communication and a fundamental mystery in neuroscience: how memory is stored over decades despite rapid turnover of proteins/RNAs. The long-term goal of the Cremins lab is to bridge chromatin folding, synaptic plasticity, and neurophysiology to elucidate how the genome’s structure-function relationship influences synaptic defects in intractable neurodevelopmental, neuropsychiatric, and neurodegenerative disorders.
-
Harshini Chandrashekar wins the Solomon R. Pollack Award for Excellence in Bioengineering Research award – Department of Bioengineering 2025. Congratulations Harshini!
-
Han-Seul Ryu presents MSTP grand rounds entitled ‘Babies, biomarkers, and the Philadelphia Tooth Fairy Project: exploring the consequences of lead poisoning across lifespan’
-
Kenneth Pham gave a great talk on his work to the UPenn Genetics, Epigenetics, and Gene expression communities on his work in heterochromatin mosaicism in fragile X syndrome.
-
Congratulations to MSTP student Han-Seul Ryu for the acceptance of her collaborative paper with the Anguera Lab entitled “B cell stimulation changes the structure and higher-order organization of the inactive X chromosome” in Cell Reports. Amazing job Han-Seul!
-
Congratulations to PhD student Rohan Patel for the acceptance of his paper entitled ” FISHnet: Detecting chromatin domains in single-cell sequential Oligopaints imaging data” in Nature Methods. https://www.biorxiv.org/content/10.1101/2024.06.18.599627v1
-
Congratulations to clinical fellow and postdoc alum Michael Guo on the publication of his paper entitled ‘Polygenic burden of short tandem repeat expansions promotes risk for Alzheimer’s disease’ in Nature Communications. https://www.nature.com/articles/s41467-025-56400-0
-
Sweety Meel joins us from Dimple Notani’s lab (https://swati.nipgr.ac.in/Scientist/SWEETY%20MEEL) as a new postdoctoral research fellow. Welcome Sweety!
-
Javed Chitaman joins us from Florida State as a new postdoctoral research fellow. Welcome Javed!
-
Skenlanda Marseille joins our lab full time for a post-bacc internship to continue her long-time work in our SEEDs program. Welcome, Skenlanda!
-
Congrats to our alum Dr. Jonathan Beagan who has now joined Precede Bio as a computational scientist. He presented his work in a poster and panel at SABCS to ascertain if chromatin signatures can be detected in liquid biopsy and used to diagnose intractable breast cancer. https://www.instagram.com/p/DD4zTogxEqK/
-
Congratulations to our alum Dr. Ji Hun Kim who has started his own laboratory at KAIST and just won his first independent grant.https://www.instagram.com/p/DDkVZRaypWc/
-
Congratulations to PhD candidate Rohan Patel – he is awarded an NIH fellowship for his work to computationally explore chromatin patterns and mechanisms of loop extrusion in sequential oligopaints imaging data.https://www.instagram.com/p/DDPftAlxE86/?img_index=5

Our work is supported by the New York Stem Cell Foundation, the Alfred P. Sloan Foundation, the National Science Foundation, the National Institute of Mental Health, the National Institute of Neural Disorders and Stroke, the National Science Foundation (NSF), NIH Common fund initiatives, the Friedreich’s Ataxia Research Alliance, the Chan Zuckerberg Initiative (CZI), the Cure Huntington’s Disease Initiative (CHDI), and the NIH 4D Nucleome Common Fund Initiative.
